Seaborn.clustermap : cluster rows and columns using different metrics
 Matthew Barrera
Matthew Barrera 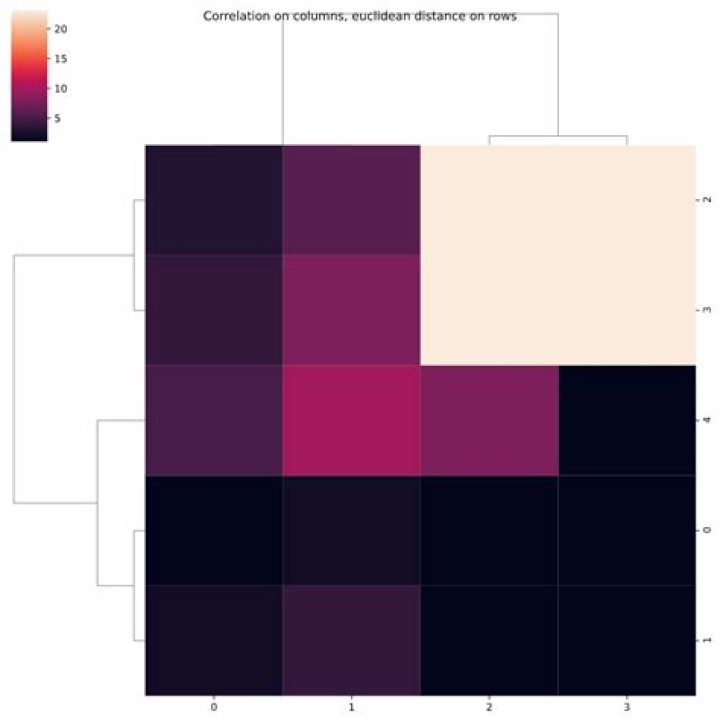
I was trying to generate a clustermap using one of the Seaborn functions. Currently, it allows me to use the same metric (Pearson, Euclidean etc.) for rows and columns, but remains difficult for using different metrics, unlike MATLAB's clustergram.
Based on this,
To use different metrics (or methods) for rows and columns, you may construct each linkage matrix yourself and provide them as {row,col}_linkage.
But does anyone know how to do that?
1 Answer
You can get the linkage matrices using scipy.cluster.hierarchy.linkage.
Here is an example:
import matplotlib.pyplot as plt
import numpy as np
import scipy.cluster
import seaborn as sns
X = np.array([[1, 2, 1, 1], [2, 4, 1, 1], [3, 6, 23, 23], [4, 8, 23, 23], [5, 10, 8, 1]])
# Clear clusters on columns by correlation: (0,1), (2,3)
# Clear clusters on rows by distance: (0,1), (2,3)
fig, axs = plt.subplots(1,2)
Z_columns = scipy.cluster.hierarchy.linkage(np.transpose(X), metric='correlation')
scipy.cluster.hierarchy.dendrogram(Z_columns, ax=axs[0])
Z_rows = scipy.cluster.hierarchy.linkage(X, metric='euclidean')
scipy.cluster.hierarchy.dendrogram(Z_rows, orientation='left', ax=axs[1])
axs[0].set_title('Columns, correlation')
axs[1].set_title('Rows, euclidean')
plt.show()# Use the computed linkage matrices in seaborn clustermap
g = sns.clustermap(X, row_linkage=Z_rows, col_linkage=Z_columns)
g.fig.suptitle('Correlation on columns, euclidean distance on rows')


